As our body develops and continues to function, our cells are constantly working. The center hub of activity is the nucleus where our genetic material – the DNA, dictates instructions for protein synthesis. Each cell has significantly more genes than it needs. Some genes (housekeeping genes) are active in all cells; they are required for the maintenance of basic cellular function, like energy production, cleaning up the cell, and so on. Other genes are needed only for ertain types of cells. To regulate genes that cells really need and genes that are not needed at present (or at all), cells use a number of processes that regulate the activity of genes, blocking genes that are not needed and opening up access to genes that the cell requires.
One of the mechanisms that helps cells achieve this complex balance is DNA methylation, the process by which chemical methyl groups attach to specific DNA nucleotides. Most often, these methyl group are attached to so called CpG sites (regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide). Like sentences that always start with capital letters, these CpG sites are often located in regions that attract special proteins to bind and start “reading” specific genes. When methyl groups are attached to these sites, the gene is “blocked” and therefore the proteins responsible for activating of this gene can’t bind there anymore. The gene is “silenced”. To activate such a “silent” gene, the methyl groups must be removed. In the first case, the genes are said to be hypermethylated, and in the second, they are hypomethylated.
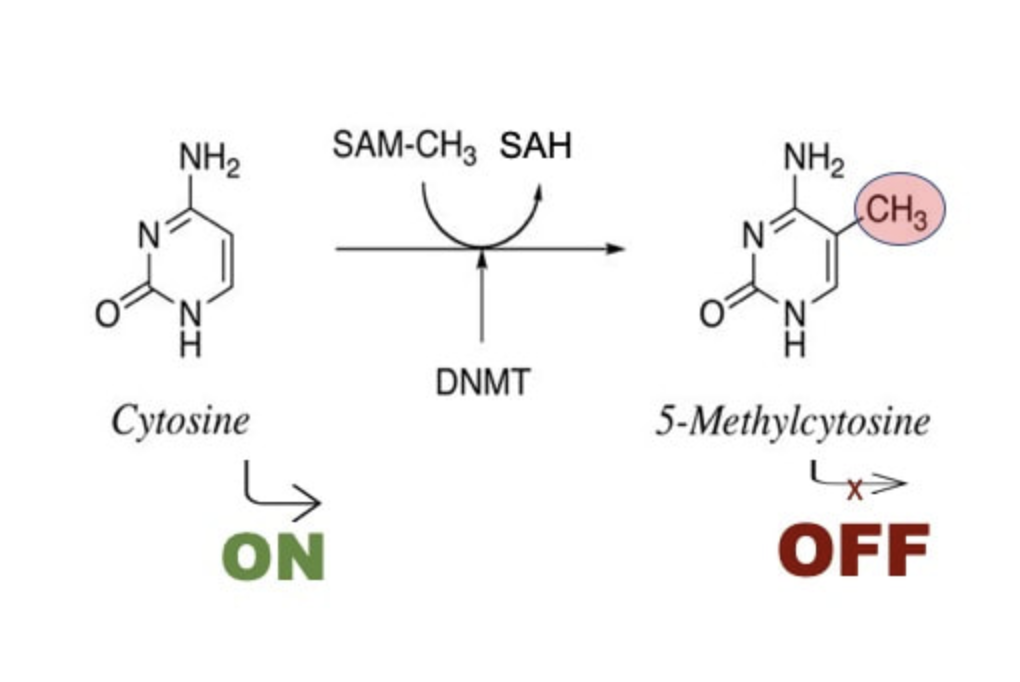

The number of methyl groups in certain genes changes with age. This discovery initiated the development of the concept of epigenetic or DNAm age clock. The researched noticed, however, that this epigenetic clock did not include muscle. However, it is important to understand how muscles age and what happens to the DNA information during this time. Our muscles are known to suffer from aging. Elderly people often suffer from so-called sarcopenia, a process during which total muscle mass is slowly reduced. Sarcopenia is one of the reasons older people become weaker over time: the muscles in the arms and legs slowly lose cells and can no longer function normally. This is one of the reasons why older people are injured so often.
Muscle cells have their own developmental program. Muscle development takes place in several stages, from the precursor cell to the full development of the muscle tissue myofibrils. At this last stage, cells can no longer divide, and new fibers can only appear with the help of “little-studied” – satellite cells. During the transformation of satellite cells into muscle cells, some genes change their methylation levels, such as MyoD, Myogenin and Six1. During aging, this complex process becomes erroneous, and although cells can divide normally, their ability to form muscle tubes is impaired.
Both the normal development of muscle tissue and the changes that occur during aging are delicate and can be influenced by many factors:
- The composition of our diet. At any stage of development, gene activity in muscle can be altered by the food we were given as children, by our current level of exercise, or by the amount of fat we accumulated as adults.
- The level of inflammation in the body. Recently, one of the inflammatory compounds in the body, TNF-α, has been shown to decrease the ability of muscle cells to regenerate. The early developmental effects of TNF-α also lead to the methylation of genes important for muscle development, MyoD, and this methylation is inherited in cells cultured for several consecutive generations. This is especially important to remember as the level of inflammation increases with age.
- On the other hand, high levels of physical activity can not only promote muscle growth, but also cause the loss of methyl groups of some genes in muscle cells.
One of the reasons that the DNA methylation age clock discovered by Steve Horvath in other tissues does not work for muscle is because methylation works differently in these cells. A study solely focused on the methylation process of muscle cells taken from young and old men showed that a completely different epigenetic clock must be used for muscle cells. Out of 500 CpG sites were found associated with epigenetic aging in the muscle tissues, and only 16 CpG sites among them were the same as Horvath’s clock.
There is one more difference. Most cells have certain genes that never active. Therefore, they are stably methylated and silent. As we age, more methylation marks are added to these “silent” genes. It seems that the system that controls them adds even more control and prevents the sudden activation of these genes. In muscle cells, this tendency of adding extra methyl tags extends to genes other than “silent” genes. What’s even more surprising is that even genes that are constantly used in muscle cells, such as genes responsible for muscle contraction, also accumulate methylation marks.
All of these facts point to a key question: It’s important to understand how the methylation process works in muscle cells. Several academic institutions, such as The Institute of Physical Performance at the Norwegian School of Sport Sciences in Oslo, The Institute for Science and Technology in Medicine (ISTM), and The School of Pharmacy and Bioengineering at Keele University at Staffordshire, UK, have collaborated and undertaken extensive research to understand the intricacies of this process. The team led by D.S. Turner and P.P. Gorski investigated methylation in muscle cells taken from young and old people, as well as in cells in culture.

Cell culture were developed from the muscle tissue, taken from young people aged 23-31 years and older people about 83 years old. They next studied DNA methylation I and gene activity in these tissues. Finally, they investigated DNA methylation after physical activity in separate groups of participants.
After the extensive research, researchers found the following:
- Muscle cells in older people had significantly more methylation marks compared to cells in younger people;
- Methylated genes in old cells have been associated with the maintenance of normal cell activity and various important biological processes;
- It is noteworthy that genes associated with muscle contractions and interactions with nerves have accumulated methyl groups;
- The group of genes with the highest methylation is associated with the development of cancer;
- The only gene that had significantly fewer methyl labels compared to younger cells coding for a special type of RNA called long non-coding RNA. This RNA is not used to build proteins. Instead, such lncRNAs are actively involved in the regulation of the activity of other genes.
When the scientists cultured muscle cells, and analyzed the patterns methylation on the genes, they have found out that:
- The cells of the elderly, as well as of the young, can be transformed into ordinary muscle myofibrils. However, at the end of the experiment, the number of myofibrils in the culture of “elderly” cells was less than in the culture of young cells;
- Older cells had lower activity of myoD and myogenin genes, which are critical for the development of myofibrils;
- At the beginning of cell culture, methylation marks were similar in both types of cells, and they changed over time, reaching the greatest differences on the 7th day of old cell culture. On the 7th day of culturing cells taken from the elderly, more than 2 thousand new methylation marks appeared, which were absent in the cultured young muscle cells. The researchers hypothesized that this is due to the fact that the methylation processes in old cells during differentiation are disrupted due to age;
- It is noteworthy that the genes that accumulated the greatest number of methylation marks were involved in important processes such as structural development of the cell, interaction with nerve cells, and other functions necessary for the normal specific functions of muscle cells;
- The 7th day turned out to be crucial for the aged cells – they had about 1500 DNA methylation position to be hypomethylated points with fewer methyl labels and 956 hypermethylated;
- Scientists discovered 5 genes that were equally affected both in cells taken directly from tissue cells and in cells that were cultured: KIF15, DYRK2, FHL2, MRPS33, ABCA17P. All of them are involved in various cell maintenance processes.
Among the groups of genes that are affected by changes in methylation in senescent cells, the HOX genes have been of particular interest. The HOX genes are critical in the development of all living organisms. They determine the structure of the body and the location of various parts of the body during the development of the embryo. Some of these genes are active in muscle tissue because they direct limb development, among other processes. The HOX genes are also responsible for controlling various aspects of gene regulation and are also part of epigenetic machinery.
In the case of the HOX genes, mainly those belonging to the HOXC9-HOX13 and HOXD group have markedly higher methylation. These groups of genes are known to be involved in limb formation, and their methylation plays a particularly important role in various types of muscle tissue. Despite the effect of methylation on gene activity, the activity of only three HOX genes changed – HOXB1, HOXC1-AS3, HOXA3. Interestingly, among these three genes, HOXB1 and HOXA3 had increased methylation and were less active, while HOX-AS3 (long non-coding RNA) had less methylation and was more active compared to younger cells. Changes in the methylation of these genes may be significant, as previous research has shown that blocking the HOX-AS3 gene prevents bone loss by increasing HOXC10 levels. Another HOX gene, HOXB1, has the potential to be suppressed by inflammation, which may explain the increased methylation of this gene in the current study.
Interestingly, the HOX genes, including HOXB1, HOXA3, HOXD12, and HOC4, which accumulate methylation during aging, were demethylated in people who actively engaged in physical activity. Likewise, groups of genes critical for cell-to-cell communication and normal cell development also lost DNA methylation after physical activity. Based on these findings, it is possible that physical activity may have an “anti-aging” effect.
This study is important because it looks closely at how aging affects muscle cells. Since the methylation of genes in these cells is partially reversible – as we can see in the people who actively engaged in physical activity – it is possible that by understanding how muscle cells regulate their genes as we age, we can reverse unfavorable processes in older people.
References
- Horvath, S. [December, 2013]. DNA methylation age of human tissues and cell types. Genome Biology, 14 Article № 3156. Retrieved November 11, 2020 from: https://genomebiology.biomedcentral.com/articles/10.1186/gb-2013-14-10-r115
- Santilli V., et al. [Sep-Dec, 2014]. Clinical definition of sarcopenia. Clin.Cases Miner Bone Metab., V. 11, №3, pp. 177-80.
- Sharples, A. P., C. E. Stewart, and R. A. Seaborne [August, 2016]. Does skeletal muscle have an ‘epi’‐memory? The role of epigenetics in nutritional programming, metabolic disease, aging and exercise. Aging Cell, 15, №5, pp. 603-616.
- Maples, J.M. et al. [May 2015]. Lipid exposure elicits differential responses in gene expression and DNA methylation in primary human skeletal muscle cells from severely obese women. Physiol. Genomics, V. 47,№ 5, pp. 139-146
- Sharples, A. P. et al. [June, 2016]. Skeletal muscle cells possess a ‘memory’ of acute early life TNF-α exposure: role of epigenetic adaptation. Biogerontology, V. 17, №3,pp.603-17
- Yin, H., F. Price, and M. A. Rudnicki [January, 2013]. Satellite cells and muscle stem cell niche. Physiol. Rev., V. 93, №1, pp.23-67
- Zykovich, A. et al. [April, 2014]. Genome-wide DNA methylation changes with age in disease-free human skeletal muscle. Aging Cell, V.13, №2, pp. 360-366
- Myers, P.Z. (2008). Hox genes in development: the Hox Code. Nature Education 1(1):2. Retrieved November 18, 2020 from: http://scienceblogs.com/pharyngula/2007/09/the_hox_code.php
- Tsumagari, K. et al. [August, 2013]. DNA methylation and differentiation: HOX genes in muscle cells. Epigenetics and Chromatin, V. 6, Article number: 25. Retrieved November 18, 2020 from: https://epigeneticsandchromatin.biomedcentral.com/articles/10.1186/1756-8935-6-25
- Turner D.C. et al (2020) DNA methylation across the genome in aged human skeletal muscle tissue and muscle-derived cells: the role of HOX genes and physical activity https://www.nature.com/articles/s41598-020-72730-z